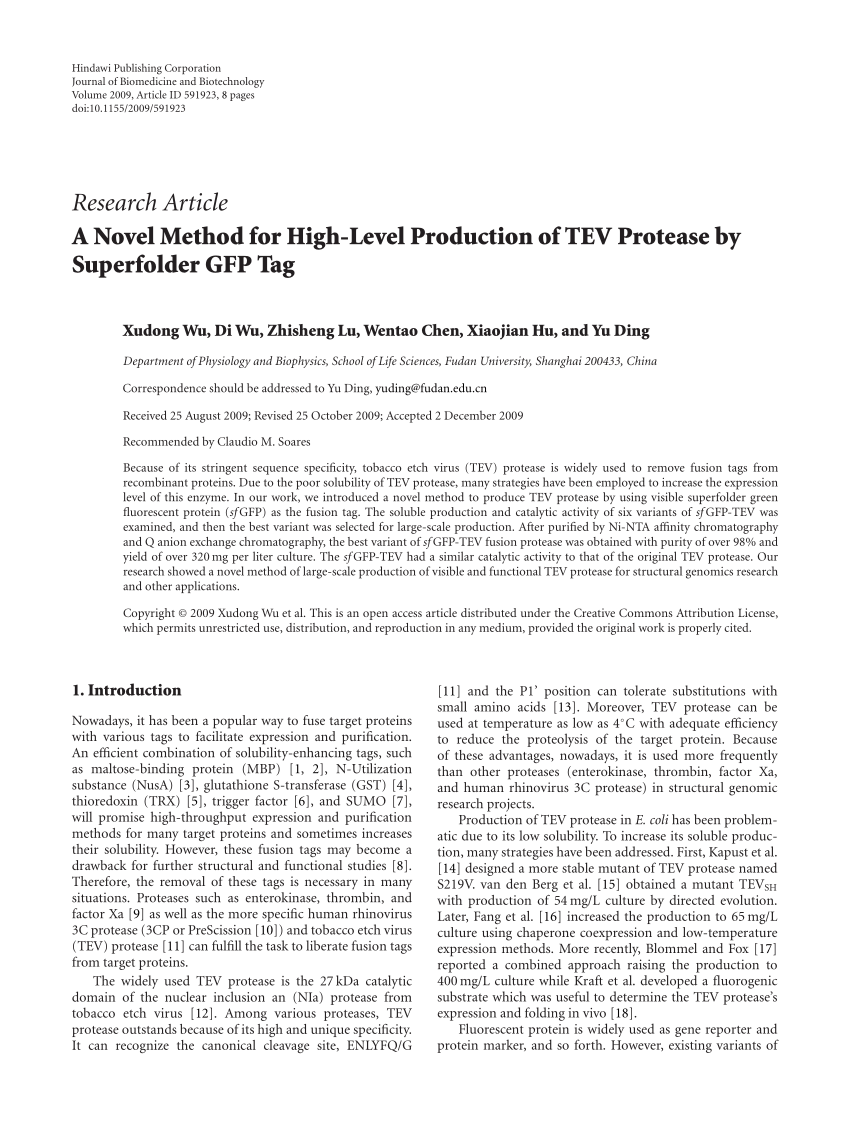
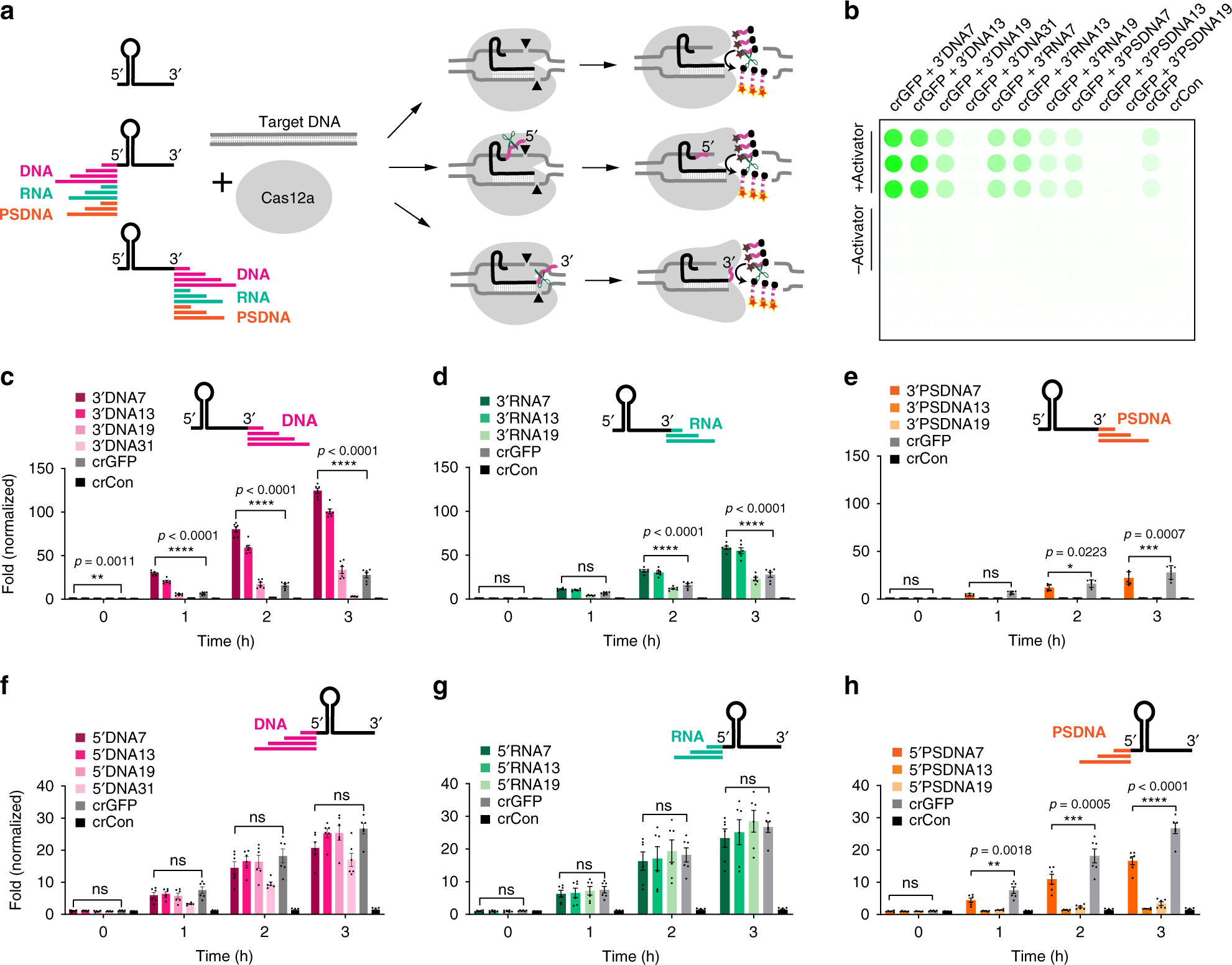
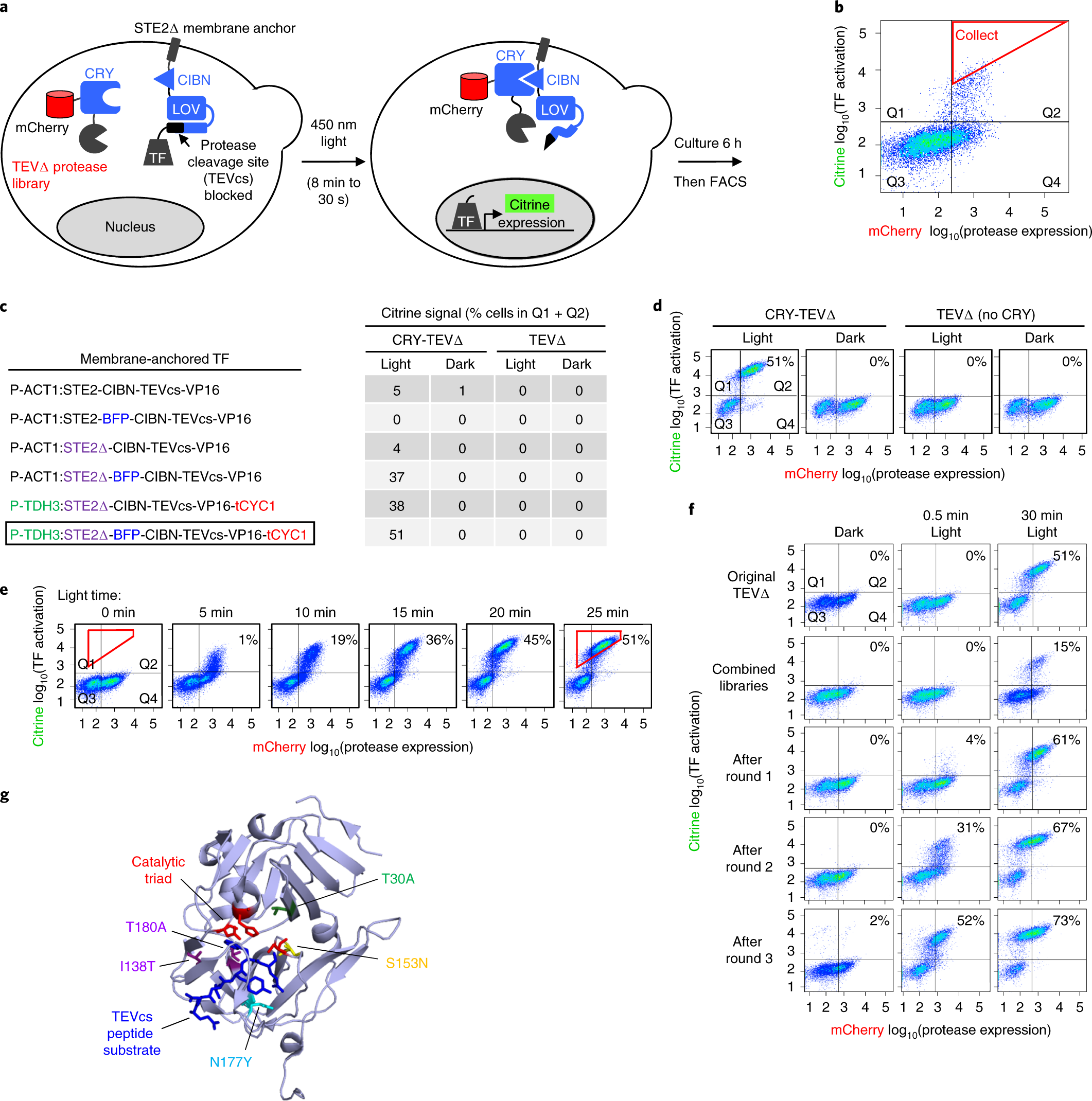
Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries
Tobacco etch virus (TEV) protease with multiple mutations to improve solubility and reduce selfв•'cleavage exhibits enhance
Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries
Tobacco etch virus (TEV) protease with multiple mutations to improve solubility and reduce selfв•'cleavage exhibits enhance
Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries

Improved yield, stability, and cleavage reaction of a novel tobacco etch virus protease mutant | SpringerLink

What conclusion I should draw with higher Km and Higher Kcat here in an enzyme kinetics data ? | ResearchGate

Phage-Related Ribosomal Protease (Prp) of Staphylococcus aureus: In Vitro Michaelis–Menten Kinetics, Screening for Inhibitors, and Crystal Structure of a Covalent Inhibition Product Complex | Biochemistry

YESS 2.0, a Tunable Platform for Enzyme Evolution, Yields Highly Active TEV Protease Variants | ACS Synthetic Biology
Tobacco etch virus (TEV) protease with multiple mutations to improve solubility and reduce selfв•'cleavage exhibits enhance
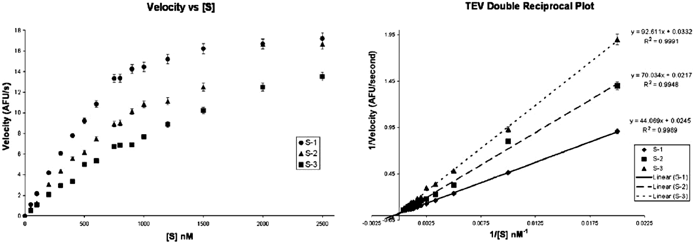
Characterization of a randomized FRET library for protease specificity determination - Molecular BioSystems (RSC Publishing) DOI:10.1039/B709290C

Quantitative Analysis of the Substrate Specificity of Human Rhinovirus 3C Protease and Exploration of Its Substrate Recognition Mechanisms | ACS Chemical Biology

Proteolytic Cleavage—Mechanisms, Function, and “Omic” Approaches for a Near-Ubiquitous Posttranslational Modification | Chemical Reviews
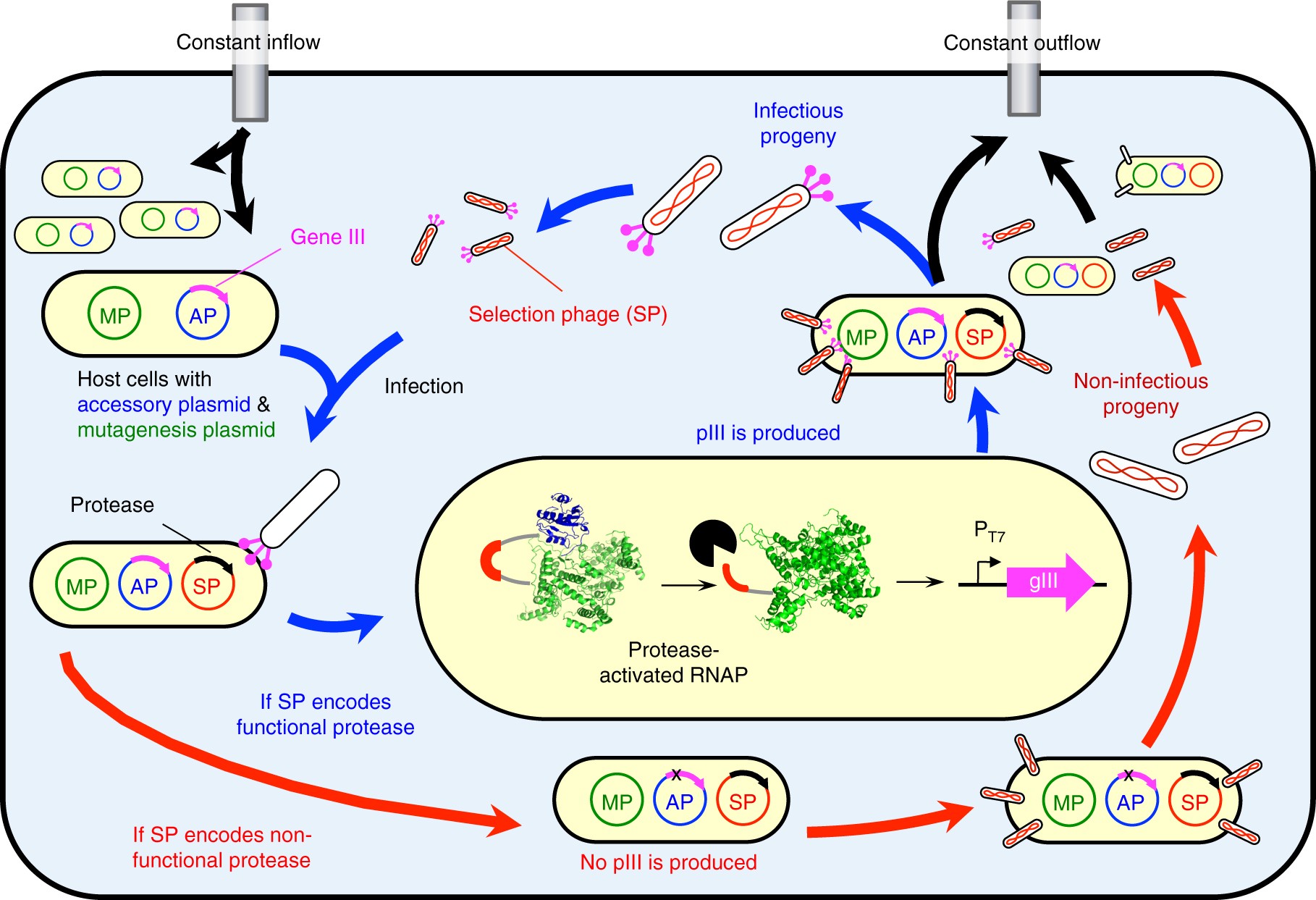
Phage-assisted continuous evolution of proteases with altered substrate specificity | Nature Communications
Engineering of TEV protease variants by yeast ER sequestration screening (YESS) of combinatorial libraries

YESS 2.0, a Tunable Platform for Enzyme Evolution, Yields Highly Active TEV Protease Variants | ACS Synthetic Biology